Calculating advanced aggregations with Elastic Search and Django
Refine.bio is a Django project that uses elastic search to power a webpage, that allows people to search transcriptome data in several experiments. In the beginning of the project we started using regular postgresql queries until the performance got shitty.
It’s relatively straightforward to use django-elasticsearch-dsl-drf to index a set of objects in elastic search and serve them through an api endpoint. However things get more complicated if you need something less conventional. For example, in our case we allow people to search for experiments, each one associated with multiple samples through a many-to-many relationship. We wanted to count the number of unique Samples for each one of the filters returned. Something that quickly pops to the eye, are elastic search’s aggregations to perform computations on the indexed documents.
The final solution consisted on indexing all sample ids for each experiment and then using the cardinality aggregation to estimate the number of unique samples for each filter category. The real challenge came down to knowing how to implement that into the project, since there’re several packages built on top of each other that each adds a different layer of abstraction.
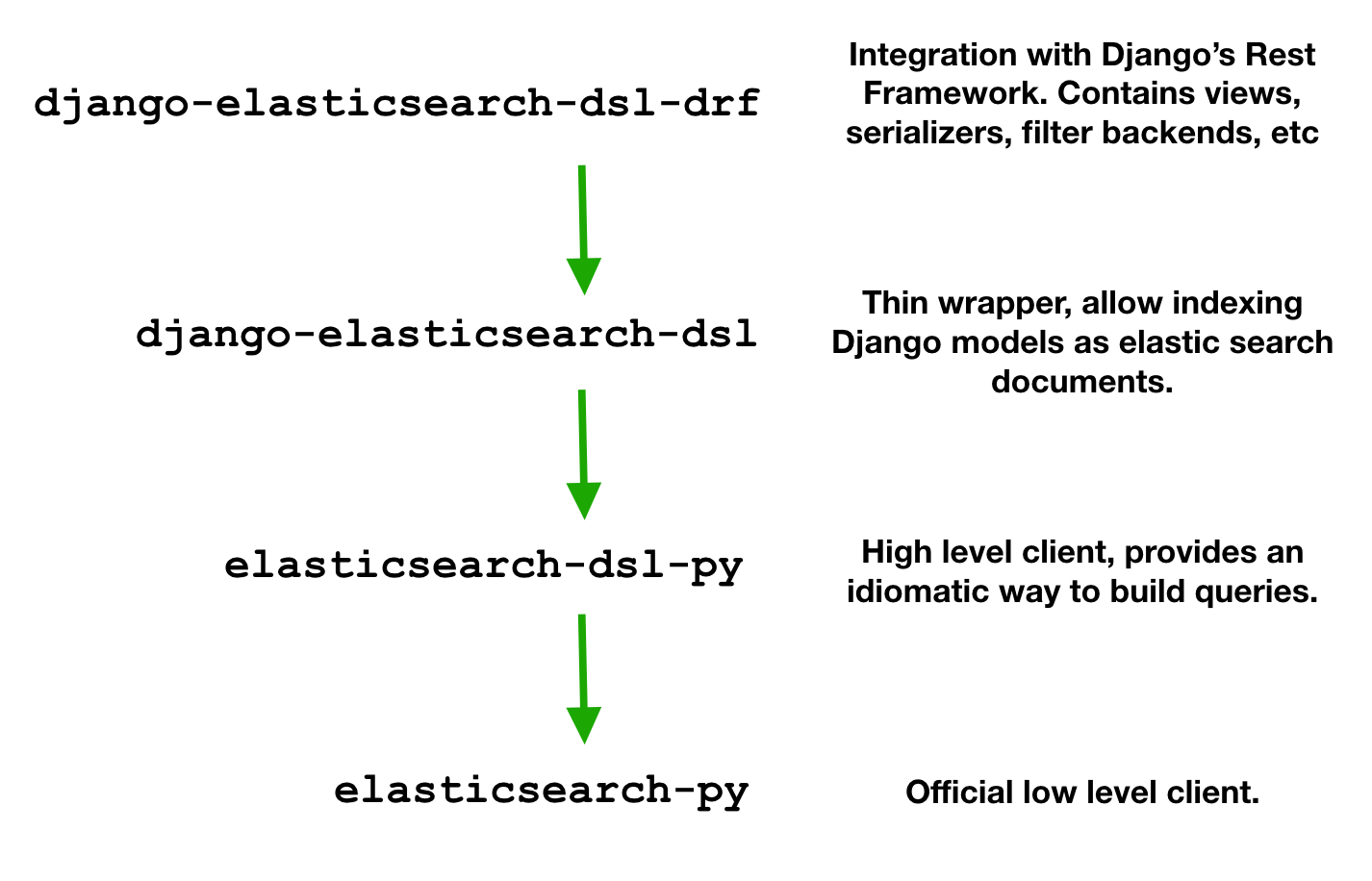
The top level package django-elasticsearch-dsl-drf “provides views, serializers, filter backends, pagination and other handy add-ons”, basically everything you need to integrate elastic search with Django’s Rest Framework. It’s built on top of django-elasticsearch-dsl and requires the document object to be defined by it.
Elastic search document objects contains specifications of how a given model should be serialized and indexed. django-elasticsearch-dsl provides helpers for this, however it’s only a “thin wrapper” around elasticsearch-dsl-py, a “high-level library whose aim is to help with writing and running queries against Elasticsearch”.
All those packages depend on the official low-level client elasticsearch-py. This package is completely abstracted by the others, and I didn’t need to use any objects from it. Mentioning here for completeness.
The implementation
As mentioned before the implementation consisted of two steps. First getting the Sample ids indexed for each document
class Experiment(models.Model):
# ...
@property
def downloadable_samples():
# ...
return sample_ids
And the field on the ExperimentDocument so that it gets indexed.
class ExperimentDocument(DocType):
class Meta:
model = Experiment
# ... other fields
downloadable_samples = fields.ListField(
fields.KeywordField()
)
django-elasticsearch-dsl-drf provides a way to add nested aggregations/facets. However this works when you need to add additional computations to the set of facets, but in the case of refine.bio we needed to add additional computations to the set of filters that were being returned by FacetedSearchFilterBackend.
I couldn’t find any solution documented for this, so I looked into the code of FacetedSearchFilterBackend and noticed that the method aggregate could be extended to add additional aggregations for each bucket.
class FacetedSearchFilterBackendExtended(FacetedSearchFilterBackend):
def aggregate(self, request, queryset, view):
facets = self.construct_facets(request, view)
for field, facet in iteritems(facets):
agg = facet['facet'].get_aggregation()
queryset.aggs.bucket(field, agg)\
.metric('total_samples', 'cardinality', field='downloadable_samples', precision_threshold=40000)
return queryset
Conclusion
There are multiple packages built on top of each other that are relevant to get Elastic Search working on a Django project. It’s important to know what each one does to find the appropriate documentation, although in some cases there’s no way around checking the code directly.